SCIN: A new resource for representative dermatology images
March 19, 2024
Pooja Rao, Research Scientist, Google Research
The Skin Condition Image Network (SCIN) dataset offers a diverse and representative collection of skin condition images, bridging important gaps for AI development, medical research, and equitable healthcare tools.
Quick links
Health datasets play a crucial role in research and medical education, but it can be challenging to create a dataset that represents the real world. For example, dermatology conditions are diverse in their appearance and severity and manifest differently across skin tones. Yet, existing dermatology image datasets often lack representation of everyday conditions (like rashes, allergies and infections) and skew towards lighter skin tones. Furthermore, race and ethnicity information is frequently missing, hindering our ability to assess disparities or create solutions.
To address these limitations, we are releasing the Skin Condition Image Network (SCIN) dataset in collaboration with physicians at Stanford Medicine. We designed SCIN to reflect the broad range of concerns that people search for online, supplementing the types of conditions typically found in clinical datasets. It contains images across various skin tones and body parts, helping to ensure that future AI tools work effectively for all. We've made the SCIN dataset freely available as an open-access resource for researchers, educators, and developers, and have taken careful steps to protect contributor privacy.
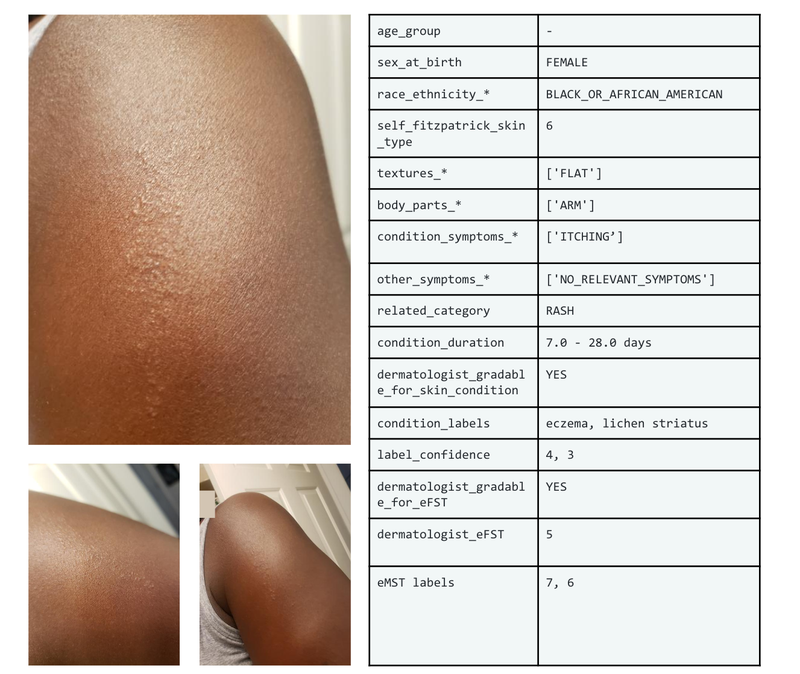
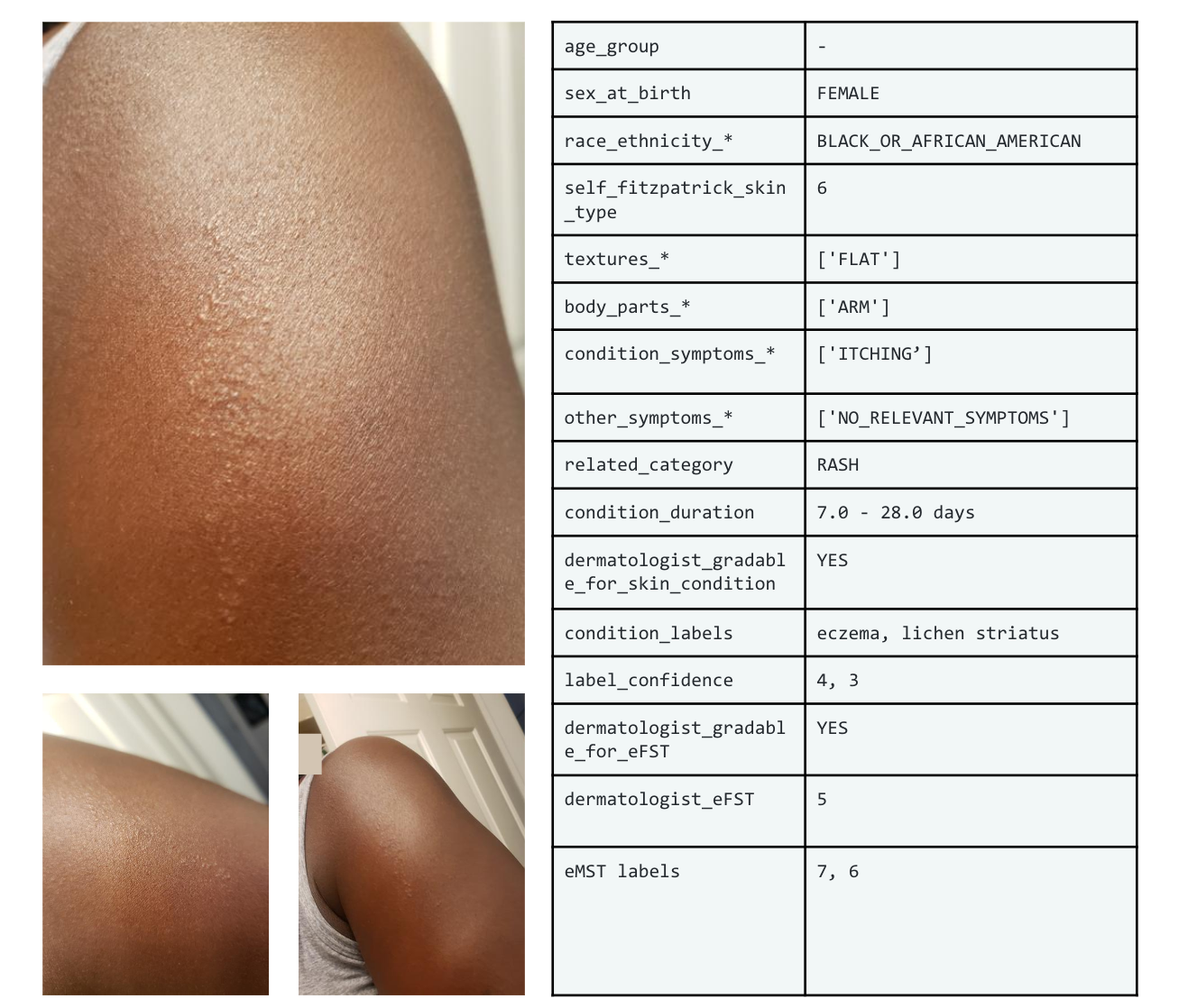
Example set of images and metadata from the SCIN dataset.
Dataset composition
The SCIN dataset currently contains over 10,000 images of skin, nail, or hair conditions, directly contributed by individuals experiencing them. All contributions were made voluntarily with informed consent by individuals in the US, under an institutional-review board approved study. To provide context for retrospective dermatologist labeling, contributors were asked to take images both close-up and from slightly further away. They were given the option to self-report demographic information and tanning propensity (self-reported Fitzpatrick Skin Type, i.e., sFST), and to describe the texture, duration and symptoms related to their concern.
One to three dermatologists labeled each contribution with up to five dermatology conditions, along with a confidence score for each label. The SCIN dataset contains these individual labels, as well as an aggregated and weighted differential diagnosis derived from them that could be useful for model testing or training. These labels were assigned retrospectively and are not equivalent to a clinical diagnosis, but they allow us to compare the distribution of dermatology conditions in the SCIN dataset with existing datasets.
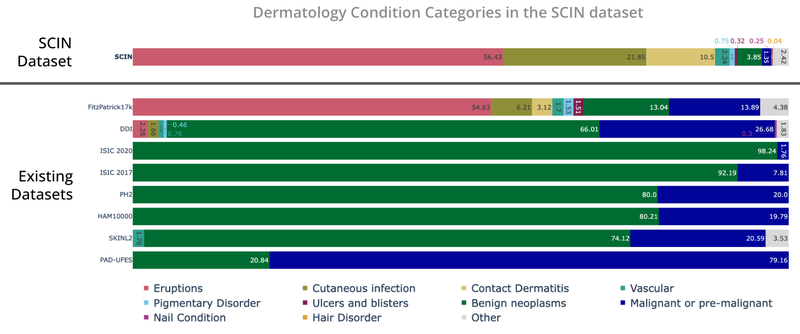
The SCIN dataset contains largely allergic, inflammatory and infectious conditions while datasets from clinical sources focus on benign and malignant neoplasms.
While many existing dermatology datasets focus on malignant and benign tumors and are intended to assist with skin cancer diagnosis, the SCIN dataset consists largely of common allergic, inflammatory, and infectious conditions. The majority of images in the SCIN dataset show early-stage concerns — more than half arose less than a week before the photo, and 30% arose less than a day before the image was taken. Conditions within this time window are seldom seen within the health system and therefore are underrepresented in existing dermatology datasets.
We also obtained dermatologist estimates of Fitzpatrick Skin Type (estimated FST or eFST) and layperson labeler estimates of Monk Skin Tone (eMST) for the images. This allowed comparison of the skin condition and skin type distributions to those in existing dermatology datasets. Although we did not selectively target any skin types or skin tones, the SCIN dataset has a balanced Fitzpatrick skin type distribution (with more of Types 3, 4, 5, and 6) compared to similar datasets from clinical sources.
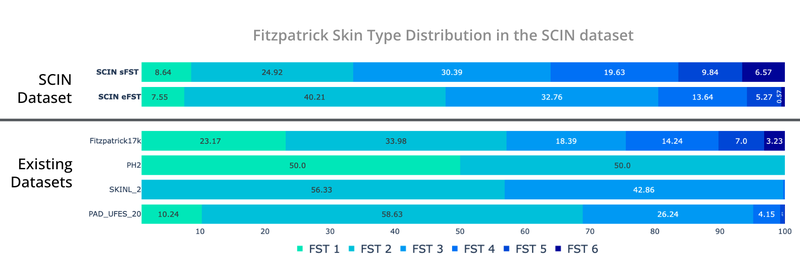
Self-reported and dermatologist-estimated Fitzpatrick Skin Type distribution in the SCIN dataset compared with existing un-enriched dermatology datasets (Fitzpatrick17k, PH², SKINL2, and PAD-UFES-20).
The Fitzpatrick Skin Type scale was originally developed as a photo-typing scale to measure the response of skin types to UV radiation, and it is widely used in dermatology research. The Monk Skin Tone scale is a newer 10-shade scale that measures skin tone rather than skin phototype, capturing more nuanced differences between the darker skin tones. While neither scale was intended for retrospective estimation using images, the inclusion of these labels is intended to enable future research into skin type and tone representation in dermatology. For example, the SCIN dataset provides an initial benchmark for the distribution of these skin types and tones in the US population.
The SCIN dataset has a high representation of women and younger individuals, likely reflecting a combination of factors. These could include differences in skin condition incidence, propensity to seek health information online, and variations in willingness to contribute to research across demographics.
Crowdsourcing method
To create the SCIN dataset, we used a novel crowdsourcing method, which we describe in the accompanying research paper co-authored with investigators at Stanford Medicine. This approach empowers individuals to play an active role in healthcare research. It allows us to reach people at earlier stages of their health concerns, potentially before they seek formal care. Crucially, this method uses advertisements on web search result pages — the starting point for many people’s health journey — to connect with participants.
Our results demonstrate that crowdsourcing can yield a high-quality dataset with a low spam rate. Over 97.5% of contributions were genuine images of skin conditions. After performing further filtering steps to exclude images that were out of scope for the SCIN dataset and to remove duplicates, we were able to release nearly 90% of the contributions received over the 8-month study period. Most images were sharp and well-exposed. Approximately half of the contributions include self-reported demographics, and 80% contain self-reported information relating to the skin condition, such as texture, duration, or other symptoms. We found that dermatologists’ ability to retrospectively assign a differential diagnosis depended more on the availability of self-reported information than on image quality.
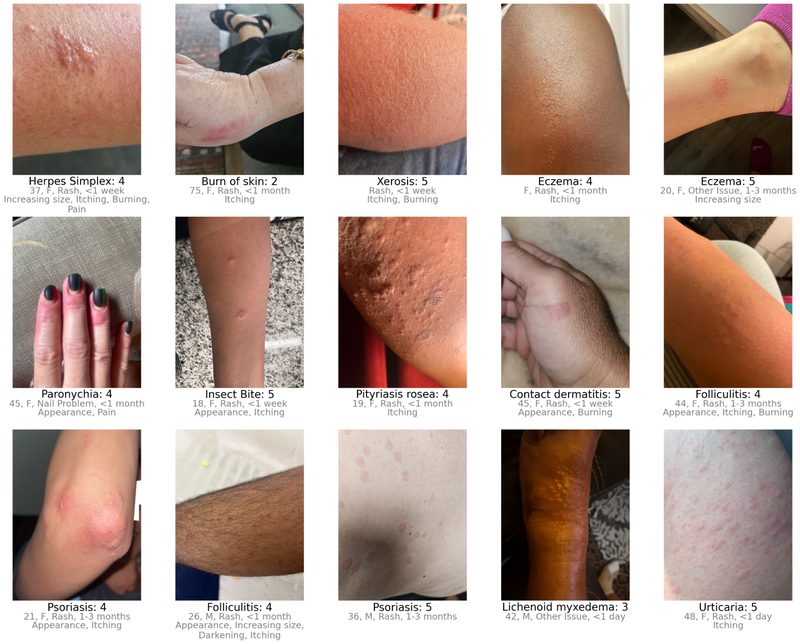
Dermatologist confidence in their labels (scale from 1-5) depended on the availability of self-reported demographic and symptom information.
While perfect image de-identification can never be guaranteed, protecting the privacy of individuals who contributed their images was a top priority when creating the SCIN dataset. Through informed consent, contributors were made aware of potential re-identification risks and advised to avoid uploading images with identifying features. Post-submission privacy protection measures included manual redaction or cropping to exclude potentially identifying areas, reverse image searches to exclude publicly available copies and metadata removal or aggregation. The SCIN Data Use License prohibits attempts to re-identify contributors.
We hope the SCIN dataset will be a helpful resource for those working to advance inclusive dermatology research, education, and AI tool development. By demonstrating an alternative to traditional dataset creation methods, SCIN paves the way for more representative datasets in areas where self-reported data or retrospective labeling is feasible.
Acknowledgements
We are grateful to all our co-authors Abbi Ward, Jimmy Li, Julie Wang, Sriram Lakshminarasimhan, Ashley Carrick, Bilson Campana, Jay Hartford, Pradeep Kumar S, Tiya Tiyasirisokchai, Sunny Virmani, Renee Wong, Yossi Matias, Greg S. Corrado, Dale R. Webster, Dawn Siegel (Stanford Medicine), Steven Lin (Stanford Medicine), Justin Ko (Stanford Medicine), Alan Karthikesalingam and Christopher Semturs. We also thank Yetunde Ibitoye, Sami Lachgar, Lisa Lehmann, Javier Perez, Margaret Ann Smith (Stanford Medicine), Rachelle Sico, Amit Talreja, Annisah Um’rani and Wayne Westerlind for their essential contributions to this work. Finally, we are grateful to Heather Cole-Lewis, Naama Hammel, Ivor Horn, Michael Howell, Yun Liu, and Eric Teasley for their insightful comments on the study design and manuscript.